郷 通子
(ごう・みちこ)
Mitiko Go
略歴
- 名古屋大学大学院理学研究科博士課程修了
- 名古屋大学理学部生物学科教授、東京大学分子細胞生物学研究所客員教授、長浜バイオ大学教授・学部長、お茶の水女子大学学長を歴任。
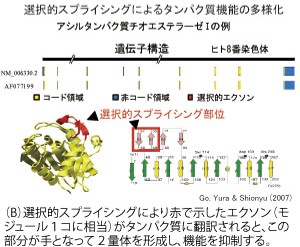
選択的スプライシングによるタンパク質の多様化
RNA編集の実態解析とその意義の重要性
モジュールに基づくタンパク質間相互作用のデザイン、モデリングとイントロンの起源など
- 研究の応用領域
- タンパク質のモジュール置換による機能創出のための独自の計算手法の、タンパク質新機能創出やケミカルバイオロジーなどへの応用。
- 産官学連携で求めるパートナー
- 「創薬のためのタンパク質モデリングと相互作用の予測」やコンピュータシミュレーションについて連携。
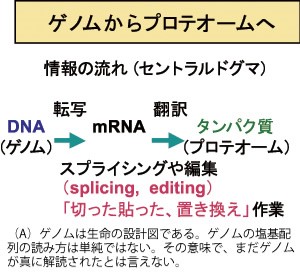
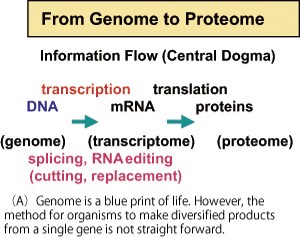
Genomic information of various species is accumulating with high speed. Translation of genomic information to proteins is not straight-forward, and there are unexpectedly complex steps to produce various kinds of proteins from a single gene (alternative splicing ) and translation to a protein after conversion of a single base of mRNA to different kind of base (RNA editing ). We analyze these biological strategies to produce diversity and aim to contribute for development of bio-technology.
Protein diversification by alternative splicing
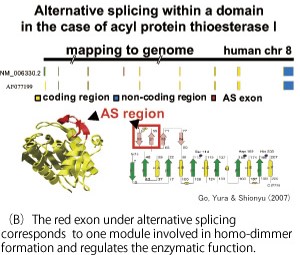
Extensive analysis of human cDNA reveals the functional role of various alternative splicing (AS). Alternative splicing is a mechanism to produce various proteins from a single gene and this is an important role of introns exiting in eukaryotic genomes. An example of AS regulating the enzymatic function is shown for acyl protein thioesterase I (Fig.(B)).
Role of RNA editing
RNA editing is a mechanism that the base sequence of a genomic DNA is not translated into a protein as it is. Enzymes edit specifically DNA bases to different bases during transcription. RNA editing sites in plant organelles are significantly skewed to residues for protein structural cores. RNA editing seems to be a mechanism to regulate protein function and expression through controlling protein folding.
Design and modeling of interaction systems based on protein modules and origin of introns, etc.
Mitiko Go, Note: Professor Nobuhiko Saitô’s contribution to statistical mechanics of biopolymers. Biophysics and Physicobiology, Special Issue “Memorial Issue for Prof. Nobuhiko Saitô” 13: 249–250 (2016).
Shionyu, M., Takahashi, K., Go, M., AS-EAST: a functional annotation tool for putative proteins encoded by alternatively spliced transcripts. Bioinformatics, 28:2076-2077 (2012).
Hijikata, A., Yura, K., Noguti, T., Go, M., Revisiting gap locations in amino acid swquence alignments and a proposal for a method to improve them by introducing solvent accessibility. Proteins: Structure, Function, and Bioinformatics, 79(6), 1868-1877 (2011).
Shionyu, M., Yamaguchi, A., Shinoda, K., Takahashi, K., Go, M., AS-ALPS: a database for analyzing the effects of alternative splicing on protein structure, interaction and network in human and mouse. Nucleic Acids Res, 37(Database issue), D305-9 (2009).
Yura, K., Go, M., Correlation between amino acid residues converted by RNA editing and functional residues in protein three-dimensional structures in plant organelles, BMC Plant Biol Jul 16; 8:79 (2008).






